Hypergraph Neural Additive Networks
In this project, we designed an interpretable hypergraph learning model for both node and hyperedge prediction tasks called HyperGraph Neural Additive Networks (HGNAN). HGNAN extends the idea of Neural Additive Models (NAMs) to hypergraph learning by integrating it with hypergraph distance-based structural information. The following figure gives an overview of our proposed model’s architecture:
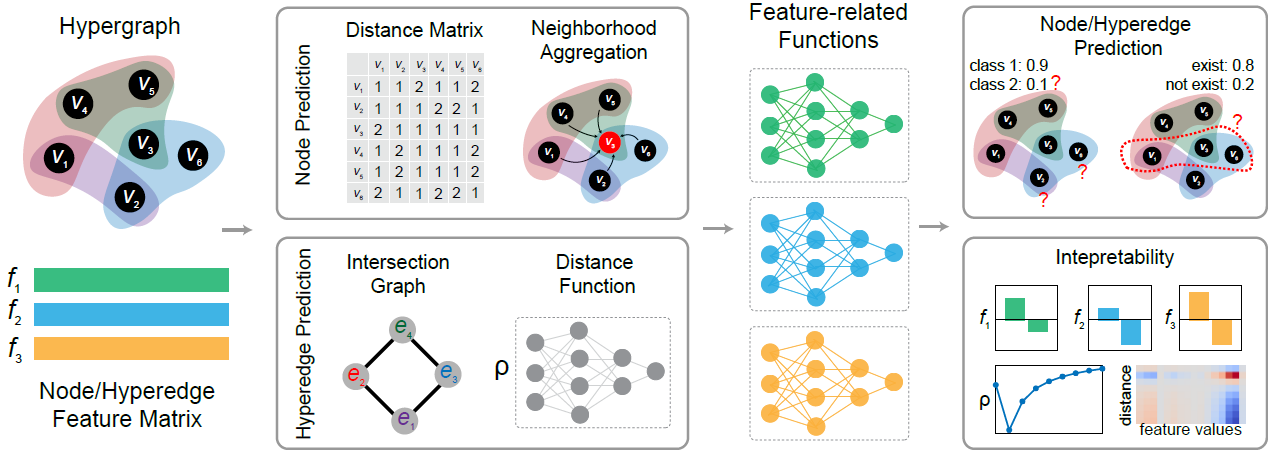
Highlights
- ✅ HGNAN can achieve on-par performance on node classification tasks while outperforming SOTA models by >10% on real-world hyperedge prediction tasks (e.g.recovering missing reactions in metabolic networks).
- ✅ HGNAN provides extra strength of providing a clear visualization of its decision making process and enabling domain experts to easily debug the model and intervene.
Method Sketch
HGNAN learns two components: a set of distance modules ${\rho_s(\cdot)}{s=1}^{s{\text{max}}}$ and a set of feature shape functions ${f_k(\cdot)}_{k=1}^{K}$. The distance module $\rho_s(\cdot)$ learns a distance function on the $s$-intersection graph, whereas each $f_k(\cdot)$ models a univariate shape for feature $k$.
The embedding for each node $i$ on the $s$-intersection graph is denoted $h_i^{(s)}$, computed by combining the $s$-level distance function with the feature shape functions. For node prediction tasks we use neighborhood-level aggregation; for hyperedge prediction tasks we use graph-level aggregation.
After obtaining $h_i^{(s)}$, the final embedding for node $i$ is a weighted sum of ${h_i^{(s)}}{s=1}^{s{\text{max}}}$ used for downstream tasks.
Results
Performance:
| Method | Zoo | Mushroom | NTU2012 | Pokec | Actor | Avg. Rank |
|---|---|---|---|---|---|---|
| MLP | 0.887 ± 0.052 | 0.965 ± 0.006 | 0.853 ± 0.012 | 0.580 ± 0.019 | 0.827 ± 0.004 | 6 |
| AllDeepSets | 0.942 ± 0.042 | 0.999 ± 0.001 | 0.876 ± 0.014 | 0.567 ± 0.008 | 0.838 ± 0.003 | 4.8 |
| AllSetTransformer | 0.973 ± 0.032 | 0.999 ± 0.001 | 0.890 ± 0.011 | 0.572 ± 0.010 | 0.836 ± 0.002 | 3.4 |
| ED-HNN | 0.950 ± 0.035 | 0.998 ± 0.002 | 0.895 ± 0.013 | 0.618 ± 0.020 | 0.856 ± 0.006 | 3.2 |
| HGNN | 0.957 ± 0.022 | 0.998 ± 0.001 | 0.872 ± 0.014 | 0.553 ± 0.014 | 0.744 ± 0.004 | 5.6 |
| HyperGCN | 0.423 ± 0.000 | 0.482 ± 0.000 | 0.796 ± 0.033 | 0.538 ± 0.014 | 0.630 ± 0.000 | 8 |
| UniGCNII | 0.950 ± 0.048 | 0.999 ± 0.001 | 0.893 ± 0.016 | 0.570 ± 0.018 | 0.828 ± 0.003 | 4.2 |
| HGNAN-node (ours) | 0.953 ± 0.030 | 0.999 ± 0.001 | 0.890 ± 0.011 | 0.634 ± 0.012 | 0.857 ± 0.004 | 3 |
| Method | iAF1260b | iJR904 | iSB619 | iYO844 |
|---|---|---|---|---|
| CHESHIRE | 0.834 ± 0.050 | 0.732 ± 0.068 | 0.730 ± 0.038 | 0.893 ± 0.047 |
| NHP | 0.732 ± 0.076 | 0.690 ± 0.090 | 0.687 ± 0.055 | 0.747 ± 0.043 |
| HyperSAGNN | 0.730 ± 0.075 | 0.753 ± 0.056 | 0.729 ± 0.162 | 0.708 ± 0.045 |
| HGNAN-edge (ours) | 0.935 ± 0.069 | 0.958 ± 0.026 | 0.977 ± 0.008 | 0.952 ± 0.181 |
Interpretability
Here we provide a small demo for visualization of HGNAN on Zoo dataset.
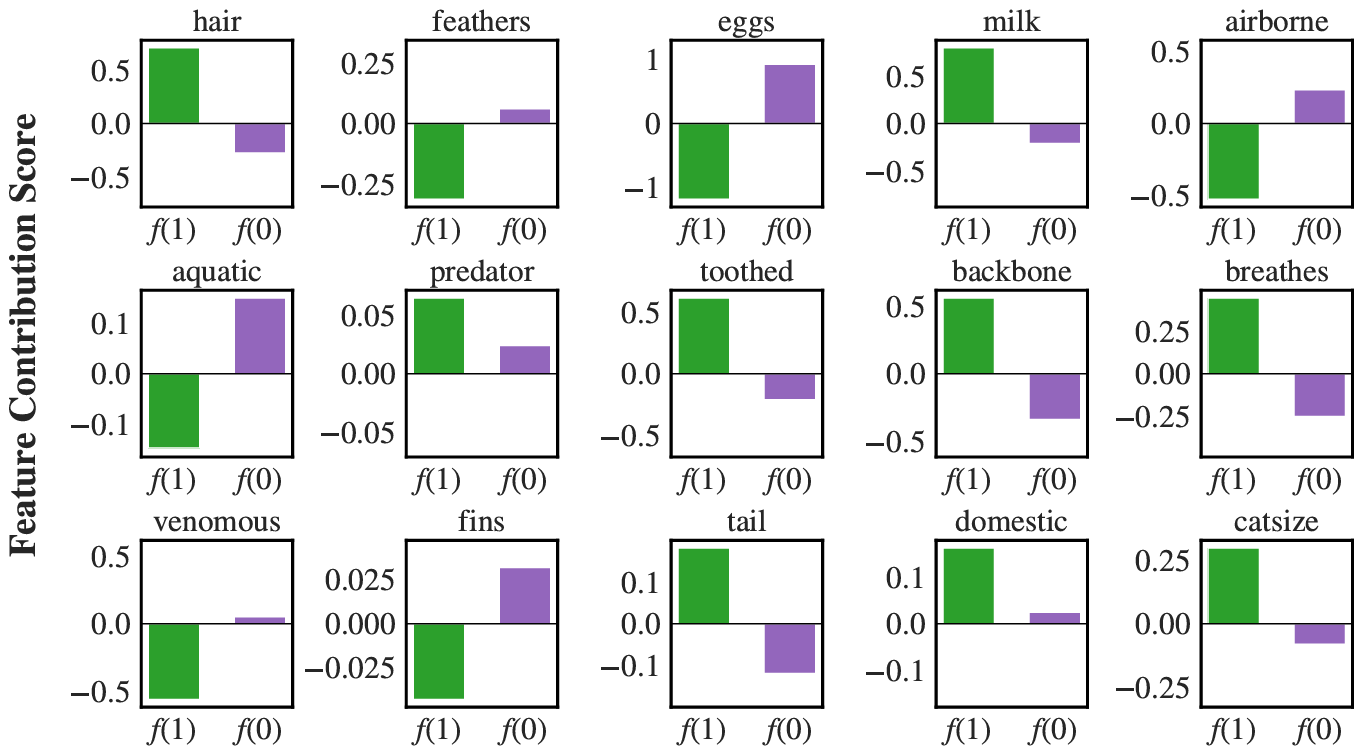
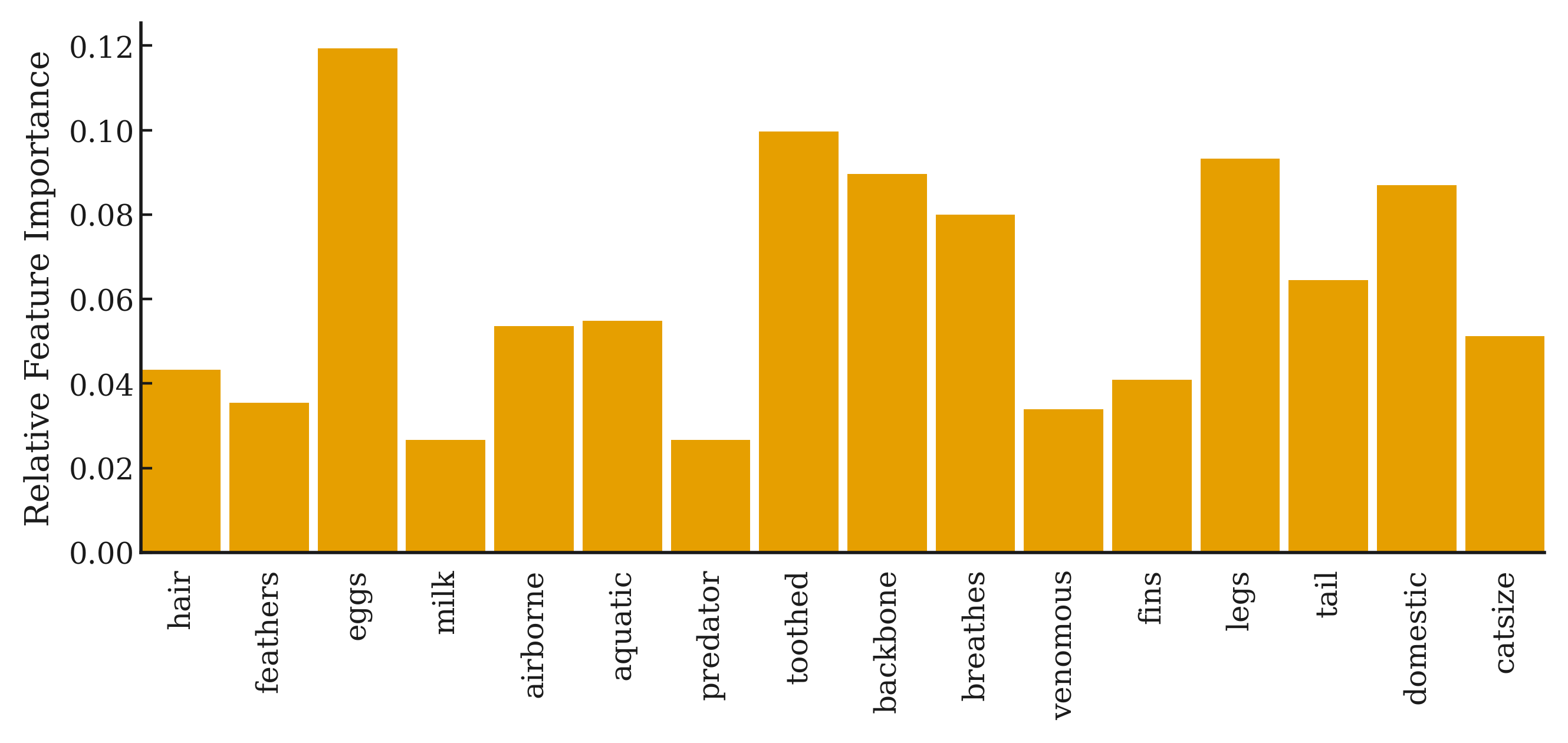
Links
- 💻 Code: GitHub